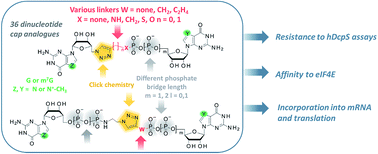
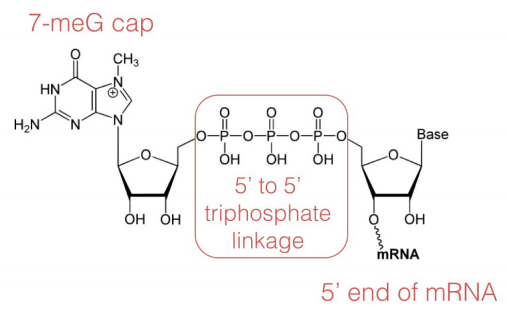
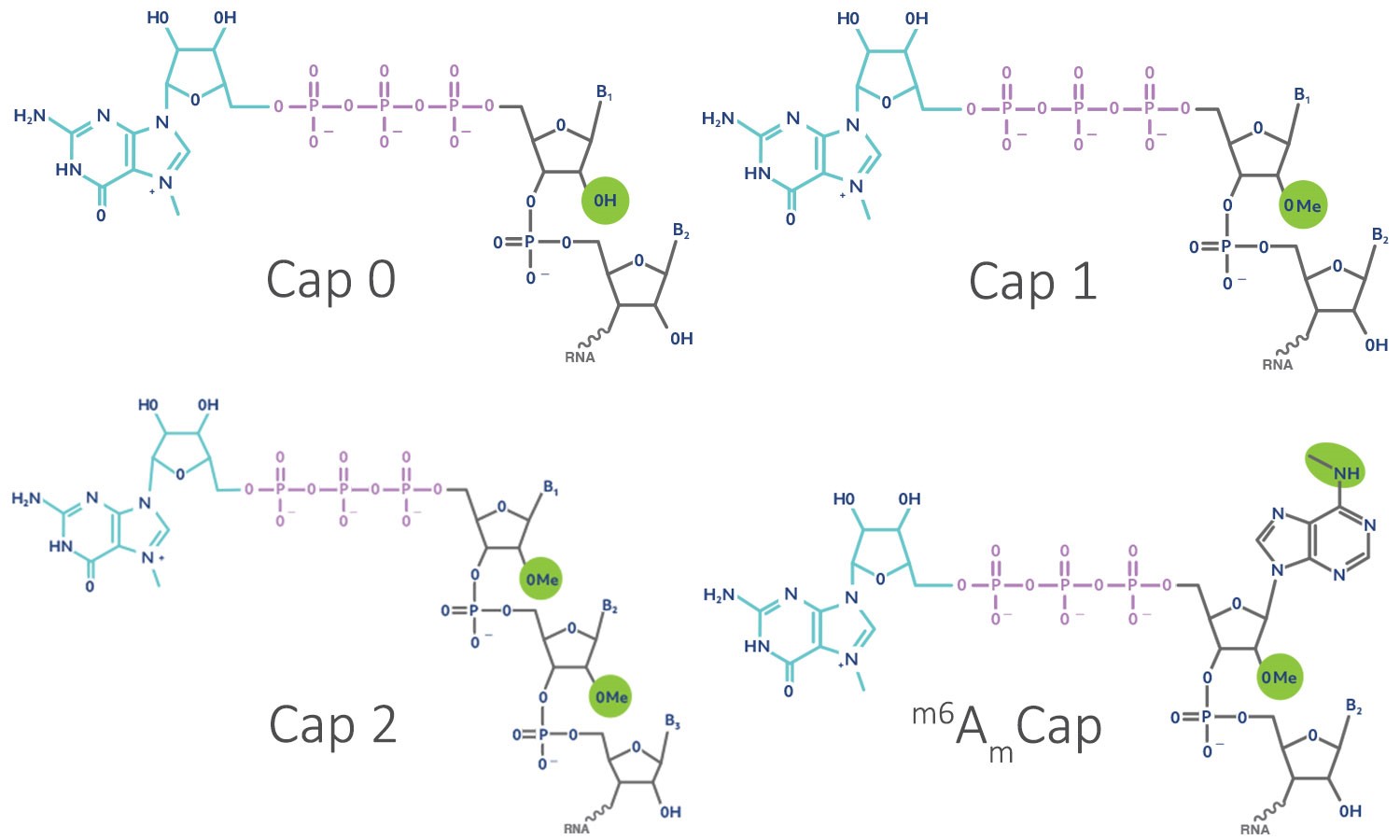
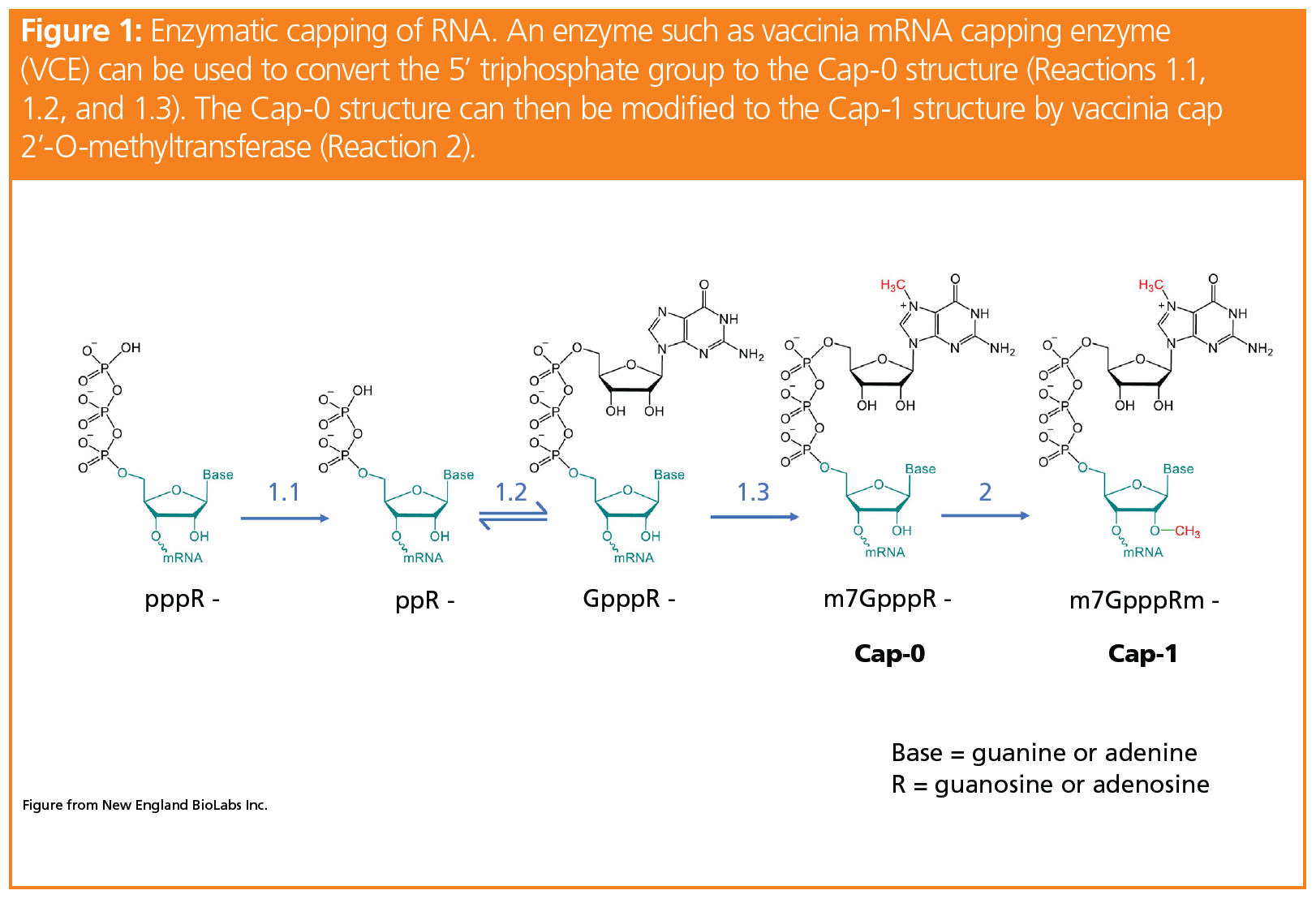
CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

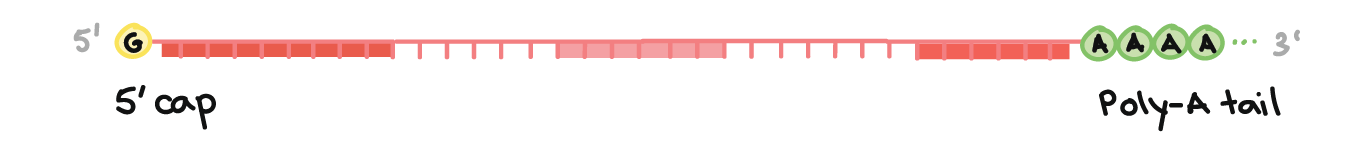
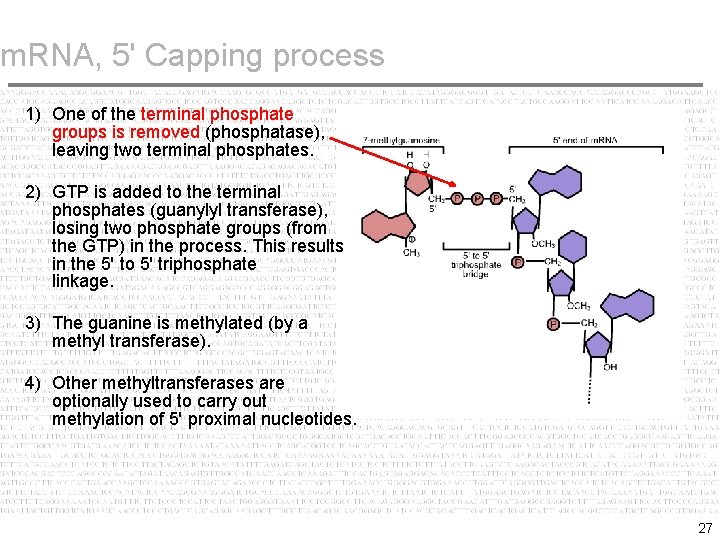
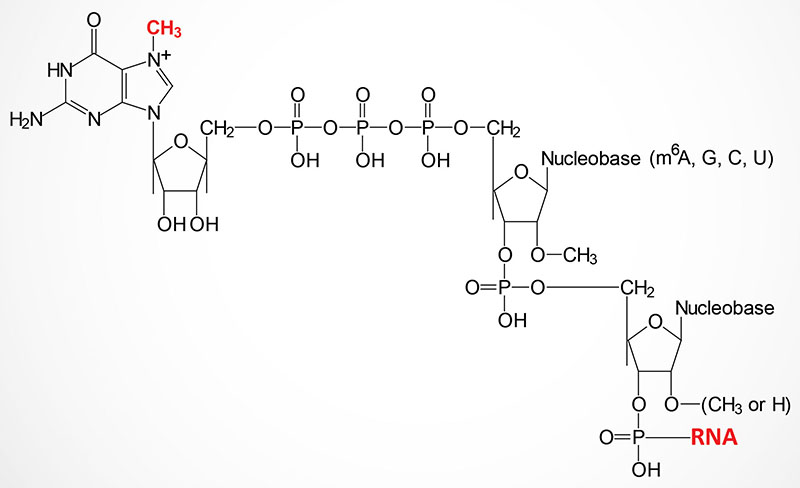
Central Dogma: Your Bio Classes - 5'-capping of processing mRNA. Capping by 7-methylguanosine. Bond form between the alpha phosphate group of guanyle compound and the beta phosphte group of the Adenosine which

5′ End Nicotinamide Adenine Dinucleotide Cap in Human Cells Promotes RNA Decay through DXO-Mediated deNADding: Cell
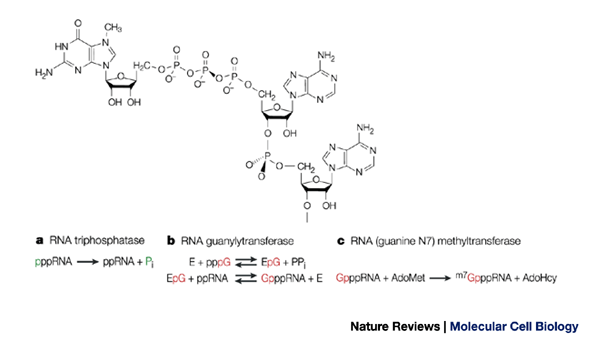
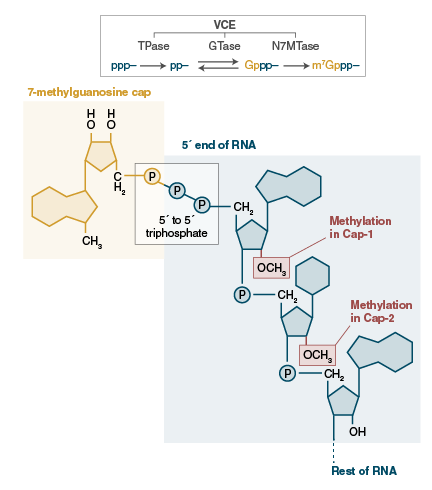
What messenger RNA capping tells us about eukaryotic evolution | Nature Reviews Molecular Cell Biology